Multi-Output Decision Trees for Lesion Segmentation in Multiple Sclerosis
Multi-Output Decision Trees for Lesion Segmentation in Multiple Sclerosis
Amod Jog, Aaron Carass, Dzung L. Pham, and Jerry L Prince
Introduction
Multiple Sclerosis (MS) is a disease of the central nervous system in which the protective myelin sheath of the neurons is damaged. MS leads to the formation of lesions, predominantly in the white matter of the brain and the spinal cord. The number and volume of lesions visible in magnetic resonance (MR) imaging (MRI) are important criteria for diagnosing and tracking the progression of MS and locating and delineating lesions manually requires tedious and expensive efforts of highly trained raters. In this paper, we propose an automated algorithm to segment lesions in MR images using multi-output decision trees.
Method
Our approach is similar to that of Germia et al. in that we also implement binary decision trees that are learned from intensity and context features, but we differ in approach. All of the images go through preprocessing. All images undergo procedures including N4 bias correction (Tustison et. al.), rigid registration to the 1mm3 MNI space (Collins et. al.), skull striping (Carass et. al.), and finally intensity standardization. When the preprocessing is done, our algorithm is made to predict the label at an entire neighborhood or patch for a given input feature vector instead of at a single voxel (The Germia et. al. approach predicts single voxels). The small patches allow us to have local context of a paticular voxel which can provide information such as the presence of lesions predominantly inside white matter. Lesion boundaries often seem to appear at slightly shifted locations in different modalities so the knowledge of predicted tissue classes in neighboring voxels is useful in determining the class of the central voxel. We then post-process the segmentations produced by our multi-output decision trees to reduce false positives.
Results
|
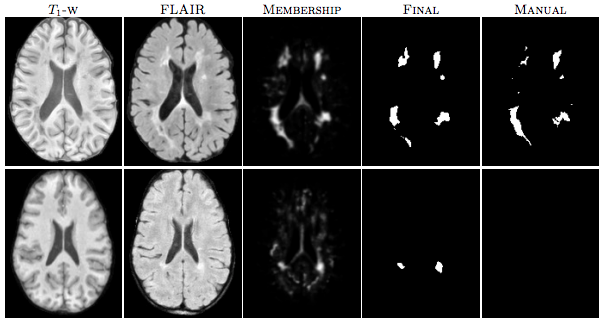
| Figure 1: The first row shows an algorithm result with a high TPR and PPV from the CHB dataset. The second row shows a low TPR and PPV result from the UNC dataset. |
We performed two experiments in order to test our algorithm. We performed the “MS Lesion Segmentation Challenge” and did an in-house MS data segmentation using an internal dataset of MS patients that was annotated by an expert rater.
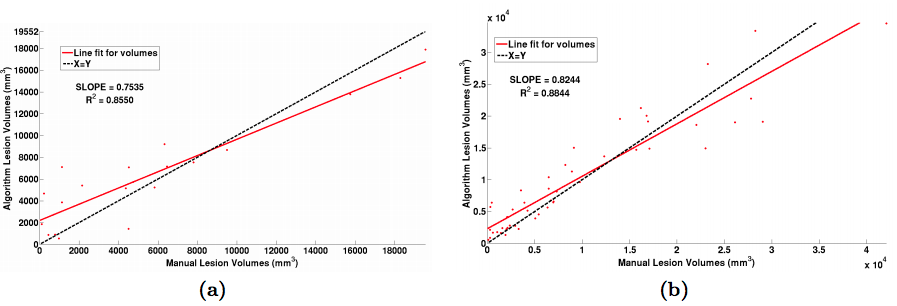
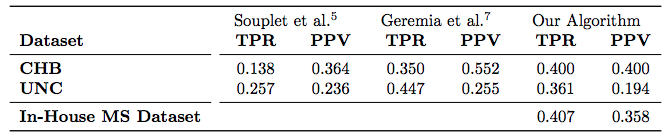
For the “MS Lesion Segmentation Challenge” we used the MSLSC Training Dataset (Styner et. al.). We compared the results with the winner of MSLSC (Souplet et. al.) and one of the current top ranked results (Germia et. al.) published after the challenge. We report some of challenge metrics and compare the estimated lesion volume with the true lesion volume. MSLSC consists of twenty subjects with T1-w, FLAIR, T2-w, and manual lesion segmentations. The data consists of twenty subjects, ten each from the University of North Carolina (UNC) center and the Children’s Hospital Boston (CHB). We randomly selected four subjects as training data and evaluated on the remainder and we also performed a leave-one-out classification of the four training data images to include them in the analysis. In the challenge, our algorithm performed generally better than the challenge winner (Souplet et. al.). Table 1 shows the results. In comparison to Germia et al., our algorithm has a better TPR-PPV balance for the CHB data and performed worse for the UNC data. The large errors in the UNC data are of the type shown in the second row of Figure 1, where the manual segmentation is debatable. Figure 2(a) shows the performance of our algorithm as a red line vs. the ideal scenario of the identity line in black. It can be seen that our algorithm slightly overestimates the lesion volume for the low lesion images while slightly underestimating for the high lesion volume images.
|
| Figure 2: (a) Volume scatter plot and linear fit for the MSLSC data and (b) our in-house MS dataset. |
|
| Table 1: Average TPR and PPV value across 10 subjects for each of the UNC and CHB datasets, reported for 3 methods. We also evaluated our algorithm on an In-House dataset of 49 subjects. |
For our second test we used an internal dataset of MS patients that was annotated by an expert rater. The in-house data consisted of 49 subjects with T1-w, T2-w and FLAIR images. T1-w images were acquired using the Magnetization Prepared Gradient Echo (MPRAGE) sequence and were acquired at a 0.82 x 0.82 x 1.1mm3 resolution. Our other scans were acquired at either 0.82 x 0.82 x 2.2mm3 for 36 subjects and at 0.82 x0.82 x 4.4mm3 for 13 subjects. Thus this dataset has considerable internal variability. We again used four random subjects as our training data and all the images were preprocessed as before in the MNI space. The results of our experiment can be seen in table 1 and in figure 2(b). Like the data we found in our first experiment, our algorithm slightly overestimates the lesion volume for low lesion load images while slightly underestimating for the high lesion load images. Figure 3 shows two subjects with high and low lesion loads and our results.
Conclusion
We have shown that our algorithm produces results which are comparable to state-of-the-art. It is computationally very fast, taking about 1-2 minutes for a 181 x 217 x 181-sized image. We have evaluated our algorithm on the training dataset of MS lesion segmentation challenge and also on our in-house collection of MS images. We would like to improve our approach by considering a different metric, which is more akin to the Gini index developed for a typical single output decision tree for purpose of classification.
Acknowledgments
This work was funded by NIH R01-NS070906, NIH/NIBIB 1R01EB017743 and NIH/NIBIB 1R21EB012756
References
- E. Germia, O. Clatz, B.H. Menze, E. Konukoglu, A. Criminisi, and N. Ayache. Spatial decision forests for MS lesion segmentation in multi-channel magnetic resonance images. NeuroImage. 57(2), 378-390, 2011.
- N. Tustison, B. Avants, P. Cook, Y. Zheng, A. Egan, P. Yushkevich, and J. Gee. N4ITK: Improved N3 bias correction. IEEE Trans. Med. Imag. 29, 1310-1320, 2010.
- D.L. Collins, C.J. Holmes, T.M. Peters, and A.C. Evans. Automatic 3-D model-based neuroanatomical segmentation. Human Brain Mapping. 3(3), 190-208, 1995.
- A. Carass, J. Cuzzocreo, M.B. Wheeler, P.-L. Bazin, S.M. Resnick, and J. L. Prince. Simple paradigm for extra-cerebral tissue removal: Algorithm and Analysis. NeuroImage. 56(4), 1982-1992, 2011.
- M. Styner, J. Lee, B. Chin, M. Chin, O. Commowick, H. Tran, S. Markovic-Plese, V. Jewells, and S. Warfield. 3D Segmentation in the clinic. A grand challenge II: MS legion segmentation. 2008.
- J. Souplet, C. Lebrun, N. Ayache, and G. Malandain. An automatic segmentation of T2-FLAIR multiple sclerosis lesions. 2008.