System for integrated neuroimaging analysis and processing of structure
System for Integrated Neuroimaging Analysis and Processing of Structure
Bennett A. Landman, John A. Bogovic, Aaron Carass, Min Chen, Snehashis Roy, Navid Shiee, Zhen Yang, Bhaskar Kishore, Dzung Pham, Pierre-Louis Bazin, Susan M. Resnick, and Jerry L. Prince
Introduction
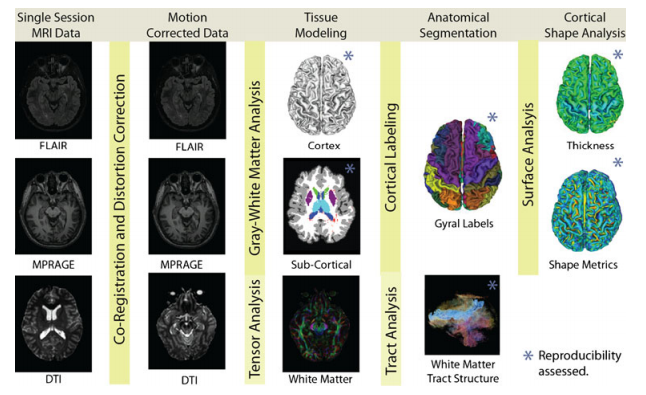
Understanding the relationship between structure and function in the brain is a crucial goal in neuroscience and imaging technology has played an important part of this research process for well over twenty years. Over the years technology has developed leading to techniques that have in turn enabled population studies using sophisticated statistical analyses from voxel based morphometry down to small world network analyses. These advances have raised the bar to look for new discoveries in ever increasing imaging data sources and ever increasing numbers of subjects and/or patients under study. Beginning now and continuing into the future, neuroscientists will require more routine analysis of increasingly larger populations with algorithms that have certified, reliable capabilities and limited need for human involvement for manual verification. This paper describes a new pipeline for multimodal analysis of neuroimaging data, the System for Integrated Neuroimaging Analysis and Processing of Structure (SINAPS).
SINAPS features several desirable properties for neuro-scientific applications. First, the algorithms are designed to take advantage of multiple MR contrasts. Second, topology preservation is obtained not only for the cerebral cortex, but for most major brain structures. This allows the results of the algorithms to be readily utilized for group shape analyses the employ high-dimensional diffeomorphic transformations. Third, a dull DW-MRI pipeline is married to this foundation by careful independent processing of DW-MRI (including motion correction of the separate gradient images) followed by deformable registration of these data to the structural data in order to remove geometric distortion caused by EPI acquisitions and magnetic susceptibility.
It is also important to note that the core algorithms have been validated individually elsewhere using comparison to manual raters, repeatability experiments, and simulations. Therefore, the accuracy against a gold standard in each of the elements of the algorithm have been (in most cases) previously determined. The objective of this study is aimed at demonstrating new analytic capabilities that became possible with integrated multimodal analysis and to provide the necessary statistical data for determination of effect size requirements in new neuroscience studies that would look to study structural and connectivity differences or change in a population.
Method
Brain Extraction: This process separates gray matter (GM) and white matter (WM) from other tissues. The primary difficulty in this task is the lack of discernible contrast between many of the tissue types. SINAPS uses the Simple Paradigm for Extra-Cerebral Tissue Removal (SPECTRE) module to extract brain from non-brain tissue by combining elastic registration with atlas-based priors, tissue segmentation and morphological techniques. SPECTRE was designed to include a thin layer of CSF surrounding the cortical surface avoiding unnecessary removal of any cortical gray matter which would adversely affect cortical reconstruction.
Brain Segmentation: Neuro-imaging research has focused on topology for the study of the cerebral cortex. We extended traditional topological constraints to describe multiple object relationships in the segmentation of brain structures using the TOpology-preserving Atlas Driven Segmentation (TOADS). TOADS maintains the relationships between all the structures to be segmented, while integrating features from tissue segmentation methods. Topological constraints prevent holes from occurring in a paticular structure and therefore suppress anomalies stemming from noise, making the method robust even without spatial regularization. What this means is that the topology is preserved during the segmentation and no topology correction is required when extracting the cortex.
Cortical Surface Estimation: Once the brain tissue has been classified into topologically correct tissue objects, locating the cortical manifold is in theory a simple matter of finding the corresponding surfaces of the cortical gray matter object. But additional processing steps lead to improved accuracy and robustness. SPECTRE, the brain extraction method, is conservative in the amount of tissue it removes. To remove additional artifacts, we developed a second-pass dura removal procedure which runs after an initial tissue segmentation. This technique also produces tissue classifications topologically consistent at the voxel-wise level. Using a feature enhancement technique and a topologically consistent deformable model, we can capture sub-voxel characteristics of the cortical manifold.
|
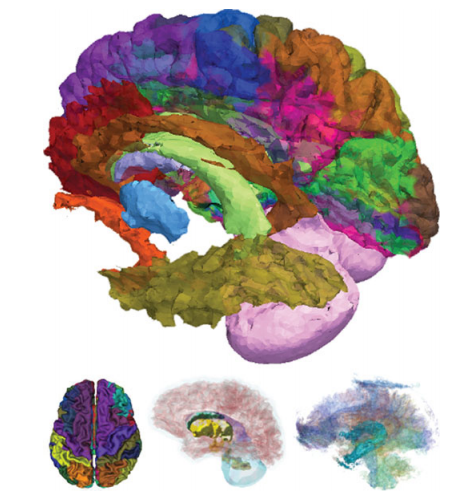
| Figure 2: Illustration of spatial fusion information (top) from surface classification (left), sub-cortical volumetry (center), and white matter tract (right) analyses |
Surface Labeling: Cortical labels corresponding to traditional neurological definitions are estimated through multi-atlas registration, label fusion, and topological correction. A collection of atlases are derived from the OASIS dataset and are non-rigidly registered to a subject’s structural image. Labels are transferred from cortical atlases through deformation of the label fields and differences between the atlas labels in subject space are reconciled using statistical fusion techniques. Five brains from the OASIS dataset with labels from the Desikan protocol were used as atlases.This technique take an intensity based approach using normalized mutual information as the similarity metric. The algorithm then constructs a deformation map as an accumulation of radial basis functions under a hierarchical schema. We account for the multiple image channels as weighted contributions within the deformable registration framework through a vectorized calculation of the difference metric. Including information from multiple channels allows the final deformation to represent anatomically variability that may only be discernible in certain image contrasts, this making the most of all available image channels. To fuse the atlas labels together on the cortical surface of the subject, we have developed an extension to the STAPLE algorithm. STAPLE estimates a true segmentation and a reliability characterization for each atlas in a collection and was originally presented on voxel level data that included a Markov random field to model spatially correlated structures.Our extended approach operates on surface labels
with a mesh-based Markov random field to account for spatial consistency of multiple labels on a mesh. After estimation, a secondary topological correction step is performed in which holes in the label definitions are filled.
Surface Assessment: We use level set subtraction to estimate cortical thickness. Cortical thickness is computed from the image volume using distance transforms from inner to the outer surface. A thickness value is assigned to each voxel in the volume between the two surfaces and is defined as the sum of the distances from the voxel to each of the two surfaces. The reconstructed cortical surface is generated in the same coordinate space as the image volume, which means that each vertex of a surface is generated in the same coordinate space as the image volume, which means that each vertex of a surface mesh can obtain image values by directly mapping into the image data in the volume. Accordingly, we obtain measures of cortical thickness at each surface vertex using trilinear interpolation applied in the image volume containing estimates of cortical thickness at volumetric grid points.
White Matter Tract Labeling: In the SINAPS pipeline, we integrate DTI and structural processing by assimilating CATNAP. To process the DW-MRI data, images are co-registered to account for motion and distortion, the gradient tables are corrected for the anatomical position relative to diffusion sensitization, and diffusion tensor sand tensor contrasts are computed. A number of strategies have been shown to be useful in correcting for these distortions and with SINAPS we use the mutual-information registration approach since it does not require additional MR image acquisitions. The undistorted anatomical MR modalities are used as registration targets to correct the geometry of the distorted EPI diffusion weighted acquisitions. This simultaneously corrects geometric distortion and co-registers the diffusion weighted images to structural images.
Results
Scan-rescan multi-modal MRI sessions on 21 healthy volunteers (no history of neurological disease, 11 males, 10 females, from twenty two to sixty six years old) were acquired from the Multi-Modal MRI Reproducibility Resource. Imaging modalities included whole head acquisitions using MPRAGE (1 x 1x 1.2mm), FLAIR (1.1 x 1.1 x 1.1mm) and DTI (2.2 x 2.2 x 2.2 mm). SINAPS was run using the three MRI modalities to extract cortical, sub-cortical, and white matter structure independently for 42 scan sessions.
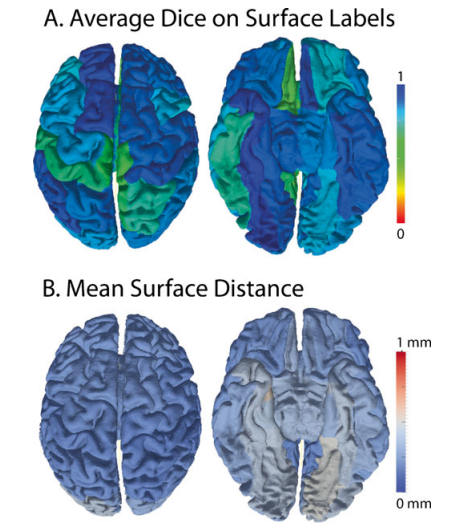
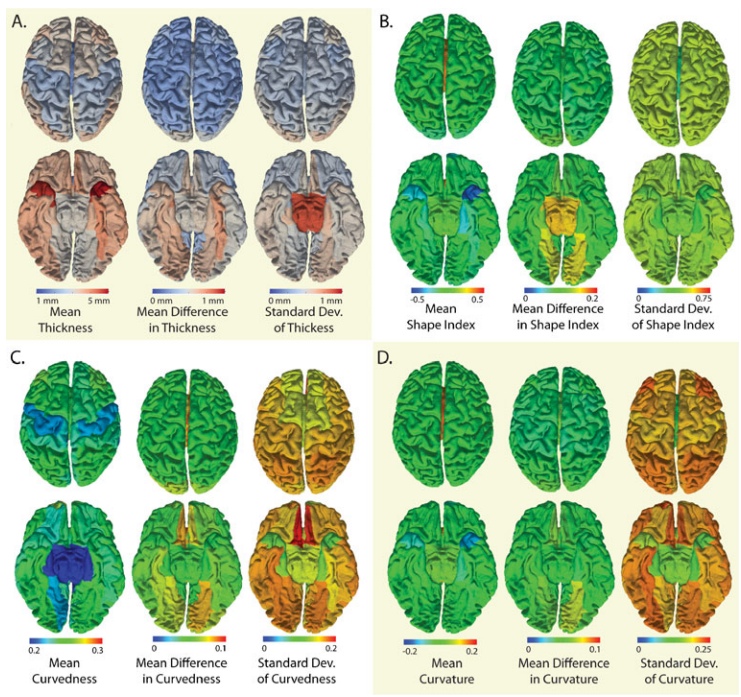
The cortical surfaces are one of the essential end-stage products of the SINAPS pipeline and are the results of multi-stage processing. The average scan-rescan Dice similarity measure between CRUISE surface labels was 0.81. To remove the confounding effects of labeling variability, the following point-wise surface statistics are reported within atlas defined regions. For atlas labeling, the surfaces were partially inflated and registered. After inter-scan registration, the average distance within a region between the estimated cortical surfaces after rigid body registration was 0.29±0.28 mm. The least reproducible regions were the entorhinal cortex and lingual gyri (0.41-0.54mm) while 44 of 40 regions exhibited less than 0.29mm surface differences. Reproducibility of cortical surface properties over all subjects are reported in Fig. 4 by the mean over gyral standard deviation of the point-wise and the standard deviation of the point-wise difference for (A) cortical Thickness, (B) shape index, (C) curvedness, and (D) mean curvature. The average cortical thickness was 3.26±0.97 mm. For all metrics the mean absolute different values (center plots) were less than the within-region standard deviation of the same metric metrics (right plots)
Both cortical connectivity and sub-cortical segmentations relay on multi-modal characterization of sub-cortical tissues. Reproducibility of volumetric brain segmentations was assessed based on average Dice coefficient of rigidly registered scan-rescan data. Reproducibility of T2 lesions volumes was approximately 0.5 Dice and reflective of the high surface/volume ration of the identified lesions and the task difficulty of distinguishing low contrast lesion boundaries. Note that the reproducibility of the remaining structures remained high despite the variability in lesion classification in necessarily reflected in label exchange with other structures. Dice overlap in the lesion areas are particularly low as this is a healthy population. One explanation for the lower Dice overlap measures is that these white matter tracts are of similar size or smaller than sub-cortical structures and more elongated. Alternatively, accuracy may be lower given the difficulty of the task.
Conclusion
SINAPS presents a consistent open-source platform which is well positioned to examine the relationship between damage to specific white matter tracts and changes in subcortical or cortical geometry defined within a specific structural network. The ability to segment white matter tracts while being robust to crossing and kissing fibers allows definition of meaningful regions of interest within the white matter, as well as the ability to automatically label tractography. The processing and analysis techniques described here have been developed for multi-modality studies, but the algorithms are modular allowing a complete analysis to be undertaken when a subset of the potential modalities are available. Each of the processing steps in the SINAPS pipeline may also be used in isolation or substituted with an algorithm with the same inputs and outputs. In conclusion, SINAPS is one viable option of many that can work with rich imaging data sets, offering diverse capabilities under a single user interface.
Acknowledgements
The Authors are appreciative of the careful feed-back from the anonymous reviewers and editor Dr. David Kennedy. This research was supported by NIH/NIA N01-AG-4-0012, NINDS5R01NS070906, IR03EB012461, 1R01NS056307, NIH/NIDAK25DA025356 (Bazin) and NIH/NINDSR01NS054255 (Pham).
Grant Support: J. L. Prince/B. A. Landman (subcontract): NIH/NIA N01-AG-4-0012, J. Prince: 1R01NS056307, B. Landman 1R03EB012461.
Publications
- B.A. Landman, J.A. Bogovic, A.Carass, M. Chen, S. Roy, N. Shiee, Z. Yang, B. Kishore, D.L. Pham, P.-L. Bazin, S.M. Resnick, and J.L. Prince, "System for Integrated Neuroimaging Analysis and Processing of Structure", Neuroinformatics, 11(1): 91-103, 2013. (doi)
- A. Carass, J.L. Cuzzocreo, M.B. Wheeler, P.-L. Bazin, S.M. Resnick, and J.L. Prince, "Simple paradigm for extra-cerebbral tissue removal: Algorithm and analysis", NeuroImage, 56(4):1982-1992, 2011. (doi)
- B.C. Lucas, J.A. Bogovic, A. Carass, P.-L. Bazin, J.L. Prince, D.L. Pham, and B.A. Landman., "The Java Image Science Toolkit (JIST) for Rapid Prototyping and Publishing of Neuroimaging Software", Neuroinformatics, 8(1):5-17, 2010. (doi)
- B.A. Landman, A.J. Huang, A. Grifford, D.S. Vikram, I.A.L. Lim, J.A.D. Farrell, J.A. Bogovic, J. Hua, M. Chen, S. Jarso, S.A. Smith, S. Joel, S. Mori, J.J. Pekar, P.B. Barker, J.L. Prince, and P.C.M. Van Zijl, "Multi-parametric neuroimaging reproducibility: a 3-T resource study", NeuroImage, 54(4):2854-2866, 2011. (doi)
- B.A. Landman, J.A. Farrell, C.K. Jones, S.A. Smith, J.L. Prince, and S. Mori, "Effects of diffusion weighting schemes on the reproducibility of DTI-derived fractional anistropy, mean diffusivity, and principal eigenvector measurments at 1.5T", NeuroImage, 36(4):1123-1138, 2007. (PubMed)
- X. Han, D.L. Pham, D. Tosun, M.E. Rettmann, C.Xu, and J.L. Prince, "CRUISE: Cortical reconstruction using implicit surface evolution", NeuroImage, 23(3):997-1012, 2004. (doi)
References
- P.-L. Bazin and D.L. Pham, "Topology-preserving tissue classification of magnetic resonance brain images", IEEE Transactions on Medical Imaging, 26(4):487-496, 2007. (PubMed)
- S.K. Warfield, K.H. Zou, and W.M. Wells, "Simultaneous truth and performance level estimation (STAPLE): and algorithm for the validation of image segmentation", IEEE Transactions on Medical Imaging, 23(7):903-921, 2004. (PubMed)
- D.S. Marcus, T.H. Wang, J. Parker, J.G. Csernansky, J.C. Morris, and R.L. Buckner, "Open Access Series of Imaging Studies (OASIS): cross-sectional MRI data in young, middle aged nondemented and demented older adults", Journal of Cognitive Neuroscience, 19(9):1498-1507, 2007. (PubMed)
Source code and binary programs developed in this paper are available via Neuroimaging Informatics Tools and Resources Clearinghouse (NITRC http://www.nitrc.org/projects/jist). The ‘DOTS WM tract segmentation project at http://www.nitrc.org/projects/dots. These tools build upon the Medical Image Processing Analysis and Visualization(MIPAV:: http://mipav.cit.nih.gov/) platform. All data are available via the Multi-Modal MRI Reproducibility Resource Resource on NITRC (http://www.nitrc.org/projects/multimoldal). The modules described herein capture the essential functionality. The details of software use and interconnections are detailed on the NITRC wiki at: http://www.nitrc.org/plugins/mwiki/index.php/jist:MultiCRUISE