
Evaluating Structural Connectomics in Relation to Different Q-space Sampling Techniques
Paulo Rodrigues1, 2, Alberto Prats-Galino3, David Gallardo-Pujol2, Pablo Villoslada4, Carles Falcon5, and Vesna Pr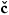 kovska4
kovska4
1Mint Labs S.L., Barcelona, Spain
2Dept. of Personality, Faculty of Psychology, UB, Barcelona, Spain
3LSNA, Facultat de Medicina, UB, Barcelona, Spain
4Center for Neuroimmunology, Department of Neurosciences, IDIBAPS, Hospital Clinic, Barcelona, Spain
5Medical Imaging Platform, IDIBAPS, Barcelona, Spain
Abstract. Brain networks are becoming forefront research in neuroscience. Network-based analysis on the functional and structural connectomes can lead to powerful imaging markers for brain diseases. However, constructing the structural connectome can be based upon different acquisition and reconstruction techniques whose information content and mutual differences has not yet been properly studied in a unified framework. The variations of the structural connectome if not properly understood can lead to dangerous conclusions when performing these type of studies. In this work we present evaluation of the structural connectome by analysing and comparing graph-based measures on real data acquired by the three most important Diffusion Weighted Imaging techniques: DTI, HARDI and DSI. We thus come to several important conclusions demonstrating that even though the different techniques demonstrate differences in the anatomy of the reconstructed fibers the respective connectomes show variations of 20%.
LNCS 8149, p. 671 ff.
lncs@springer.com
© Springer-Verlag Berlin Heidelberg 2013