 Research
Software
Research
Software
MRI
Analysis Tools
[Back/Home] [Incidental Art] [Introduction
to MRI/DTI] [Templates] [Resources]
Software Released Relating to My Research
¨ ViPAR – Medical imaging MIPAV plug-in for real-time 3D volume transformation and visualization (Java)
¨ CATNAP – Modular, cross-platform diffusion tensor imaging processing pipeline (Matlab and Java)
¨ DTI-SPOT– Cross-platform overlay tool to superimpose DTIStudio fiber tracking results on 3D volumes.
¨ Tessellation of Platonic Solids –Toolbox for representation and tessellation of platonic solids (Matlab)
¨
Philips PAR/REC File Format
Toolbox – Toolbox for reading, writing and
converting the Philips research file format to/from Analyze files (Matlab)
¨
MIPAV Philips PAR/REC File Format
(JAVA) – Native support for PAR/REC within MIPAV
and the MIPAV development environment.
¨
DTI_gradient_table_creator – Function to determine the diffusion weighting directions
relative to the voxel ordering for DTI experiments on Philips MRI scanners
(Matlab)
¨
Evaluation of DTI Methods – An open framework to evaluate tensor fitting methods
 ViPAR
ViPAR
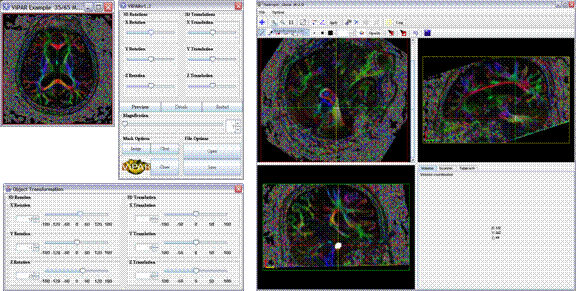
ViPAR (Visualization, Paint, Allignment and Rotation) is a MRI visualization and manipulation tool that enables real-time 3D transformation of MR volumes and delineation of regions of interest on arbitrary planes. It is available as a plugin to MIPAV.
Status: Freely available.
Platform: Java
URL: http://iacl.ece.jhu.edu/resources/
Screen shot:
 CATNAP
CATNAP
CATNAP (Coregistration, Adjustment, and Tensor-solving – a Nicely Automated Program) is an end to end data processing pipeline for Philips PAR/REC files. CATNAP performs motion correction for both diffusion and structural images using FSL FLIRT, adjusts the diffusion gradient directions for scanner settings and motion correction, and estimates tensor and derived quantities. The results are readily compatible with DTIStudio, FSL, and other tensor analysis packages.
Status: Public Release
Platform: Matlab
Screen shot:


DTI-SPOT
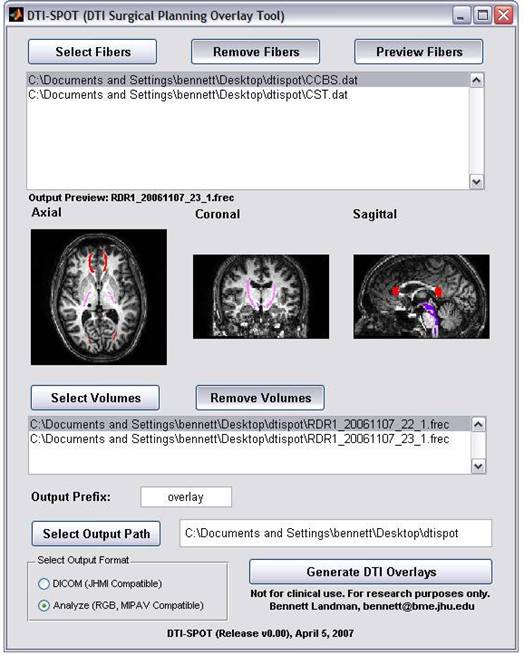
DTI-SPOT (DTI Superposition Of Tracts) is a cross-platform user interface for superimposing DTIStudio filber tracking results on 3D PAR/REC and Analyze Volumes. Combined images are available in DICOM and Analyze (RGB) format and are readily compatible with general purpose DICOM visualization packages.
Status: Internal Release
Platform: Matlab
Screen shot:
 Tessellation
of Platonic Solids
Tessellation
of Platonic Solids
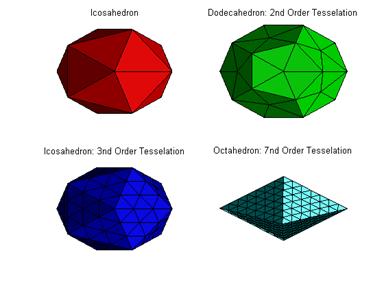
Small Matlab scripts for creating and visualizing the Platonic solids and their tessellations.
Status: Beta Release. Comments welcome. [Download]
Notes: If you find this software useful, please let me know [landman (at) jhu.edu].
Platform: Matlab (tested on v 7.0+)
Screen shot:
 Matlab
Philips PAR/REC File Format Toolbox
Matlab
Philips PAR/REC File Format Toolbox
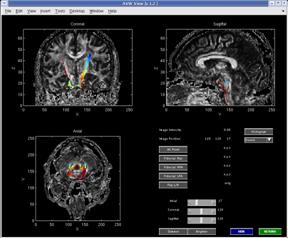
The PAR/REC file format toolbox enables robust parsing of Philips MRI research file format volumes. As of this beta release, PAR versions 3 and 4 are supported. Direct visualization is integrated with the MRI_TOOLBOX (Darren Webber). Multi-volume and arbitrary slice ordering are correctly handled. PAR/REC files may be written to or generated from Analyze format.
Functions of particular interest:
loadPARREC.m – load PAR/REC files
loadPAR.m - Parse a par header into a maltab structure
writePARRECtoAVW.m - Convert par/rec to analyze
writeAVWtoPARREC.m - Convert Analyze files to a par/rec
savePARREC.m - Save a Matlab structure to a par/rec file
savePAR.m - Save a Matlab header to a par file
Notes:
- Most scripts containing “REC” have alternate versions with “fREC”. This file format is identical to PAR/REC with the exception that the REC file is 32 bit floating point rather than 16 bit unsigned integer. The scaling defined in the PAR file is automatic.
- This toolbox supports either par/rec or PAR/REC, but both the par and rec must have the same capitalization structure.
- All par files generated by this program have “v2” appended to the end to indicate that they were not generated by a Philips scanner. We attempt to exactly preserve the format structure, but the formatting of the numbers may be different (e.g., 0.00 versus 0).
- Many other functions have been included in this toolbox that relate to various MR formats. Please let me know if you can comment on any features, bugs, or general concerns.
- Release 1.1 includes support for MIPAV’s minimal PAR/REC file format.
MIPAV Philips PAR/REC File Format (Java)
The PAR/REC file format support enables robust parsing of Philips MRI research file format volumes. PAR versions 3 and 4 are supported. Multi-volume with linear slice ordering is correctly handled. All files must be little endian.
Support has been release for internal use and will soon be available externally.
Status: See the MIPAV website (http://mipav.cit.nih.gov/) and/or the MEDIC MIPAV support forums (http://medic.rad.jhmi.edu/)
Platform: Multi-platform Java
DTI_gradient_table_creator determines the correct gradient table for a given DTI acquisition on Philips MR units. The code computes the gradient table based on minimal user input, and provides a quick and easy method to determine the correct gradient table necessary to calculate diffusion tensors (i.e., for FA, Colormaps, fiber tracking etc.)
Systematic Evaluation of Linear and Nonlinear DTI Estimation Methods: An Open Framework
A
basic premise of DTI is that the tensor formalism (e.g., assumption of Gaussian
diffusion) meaningfully represents diffusion processes, and thus the derived
contrasts are relevant. Yet, not all tensors represent physically possible
processes (e.g., those with negative eigenvalues); typical log-linear mean
squared error methods can result in these non-physical solutions. Various
non-linear tensor estimation frameworks have been developed to prevent these
problems (Tschumperlé and Deriche 2003; Cox and Glen 2006; Niethammer, Estepar
et al. 2006), while regulation and robust tensor estimation methods employ
spatial correlations to lessen the effects of noise (Mangin, Poupon et al.
2002; Chang, Jones et al. 2005). Despite the emergence of new methods, little
evidence has been presented to provide equivalent experimental comparisons or
enable well-informed selection of the appropriate tensor estimation method for
a particular task.
Last Updated: Tuesday May 27, 2008
© Copyright 2006, Bennett Landman. All rights reserved.